Thesis: 5.1 Data structures
tnet uses two basic data structures depending on the nature of the data. The first one is used to represent weighted static networks. Since most networks are sparse (i.e., the number of ties is much lower than the squared number of nodes: ), we opted for an edgelist format instead of a matrix one. This is a format that records the sender and receiver of established ties. The main advantage of this format is that it can scale to networks with many nodes as it is the number of ties, not nodes, that determine the size of the data object. Although many programmes can read edgelists, most network analysis programmes rely on an internal matrix representation, e.g. UCINET and the network-package (Barrat et al., 2004; Newman, 2001c). Conversely, Pajek, which was designed to analyse large-scale sparse networks, specifically uses an internal edgelist representation (Batagelj and Mrvar, 2007).
A binary edgelist consists of two columns that represent pairs of nodes that are tied together (e.g., the edgelist1-format in UCINET’s dl files; Borgatti et al., 2002). When a directed network is represented, the first column represents the nodes that create the ties, whereas the second column represents the target nodes. Thus, an edgelist is an matrix.
This type of list has been extended to cover weighted networks by adding a third column representing the weight of the ties (Borgatti et al., 2002). In an effort to stay consistent with existing data structures, this is also the structure used by tnet. The class of the object in R should be data.frame. This class of object allows the different columns of a table to be of different classes themselves, such as integer and numeric. Thus, the class is more efficient at storing data than a matrix which requires all columns to be numeric. The first two columns of object are assumed to be integers (i.e., the identification number of the node creating the tie and the identification number of the node receiving the tie, respectively). The third column can be real numbers (numeric) that represent the weights attached to the ties.
To illustrate the edgelist structure, we represent the directed network in Figure 16a by using the following matrix:
1 2 4 1 3 2
Table 8: Format for directed weighted edgelists: a
data.frame object in R.
To represent an undirected network, each undirected tie must be included twice — one in each direction. Therefore, the undirected network in Figure 16b should be represented by the following matrix:
1 2 4 2 1 4 1 3 2 3 1 2
Table 9: Format for undirected weighted edgelists: a
data.frame object in R.
There are a number of functions that help the users to convert other formats into the weighted edgelist format. For example, if the dataset is undirected, but there is only one entry for each tie in the edgelist, the symmetrise function adds a second entry of the edge with the identification numbers of the creator and target nodes reversed. Moreover, if the dataset is similar to an edgelist, but with only two columns (representing the identification numbers of the creator and target nodes) and multiple entries of the same tie refer to the weight of that tie (e.g., if a tie has a weight of 3, it is included three times), then the shrink_to_weighted_network-function allows the users to convert the edgelist into the correct format.
To allow for a comparison between weighted and binary network measures, the dichotomise-function creates a binary network from a weighted one. It does so by removing the ties in a weighted edgelist that fall below a certain cut-off and sets the weight to 1 for the remaining ones.
The second data structure is the longitudinal one. This structure represents network data where the sequence of ties is known, i.e. each tie is associated with a timestamp. For example, these network datasets include those obtained by studying phone and online interaction log files. In recent years, there has been a rise in the availability of this type of datasets (e.g. Ebel et al., 2002; Hall et al., 2001; Holme et al., 2004; Kossinets and Watts, 2006; Onnela et al., 2007; Panzarasa et al., 2009). This is mainly due to the fact that people communicate more through electronic mediums (Wellman, 1999), and the providers of these mediums are often required by law to keep a log of the activity where the time is explicitly stated. Yet, few methods have been developed for this type of datasets, and no general programmes have been developed to study them.
Figure 17 exemplifies the first six time steps of such a network. At a tie is created from node 1 to node 2 among the only two nodes in the network. An isolate (node 3) joins the network at
, and the tie from node 1 to node 2 is reinforced, whereas in the following time step, the tie is weakened. At
, node 1 creates a tie with the isolate node, node 3. Node 3 reciprocates this tie at
, and direct another tie towards node 2 (
). The last tie closes the non-vacuous triad starting at node 3 to node 2 through node 1 (Wasserman and Faust, 1994, 243).
The longitudinal data structure is a
data.frame object, where T is the total number of time steps. The first column represent the time at which the tie was created and should be a string with a standard Unix/SQL timestamp, i.e. YYYY-MM-DD HH:MM:SS where YYYY is the four digit year, MM is the two digit representation of the month, DD is the two digit day of the month, HH is the two digit hour of the day in 24-hour format, MM is the minutes, and SS is the seconds. The second and third column are integers that represent the identification numbers of the creator and target nodes, respectively. These numbers must follow a strict sequence, e.g. the two nodes connecting at must be node 1 and 2. An increase in this sequence reflects the inclusion of a new node. The fourth column is numeric and measures the increment to the weight of the tie. Currently, all the functions in tnet only interpret this when the incremental value is either +1 or -1. In other words, +1 refers to creation or reinforcement of the tie, whereas -1 reflects a weakening or severing of the tie. The fifth column represents the total number of nodes that have been active in the network at a given time. This column is important as it allows for inclusion of isolates in the network (i.e., nodes not represented in columns 2 and 3). Finally, nodes can join and withdraw from the network at specific times. This is signalled by a self-loop (i.e., when the second and third column have the same identification number). The fourth column controls whether it the node is joining (+1) or withdrawing (-1).
The sample network shown in Figure 17 has the following structure:
2008-09-12 13:45:00 1 2 1 2 2008-09-12 13:46:31 1 2 1 3 2008-09-12 13:49:27 1 2 -1 3 2008-09-12 13:58:14 1 3 1 3 2008-09-12 13:52:17 3 1 1 3 2008-09-12 13:54:26 3 2 1 3
Table 10: Format for longitudinal data: a
data.frame object in R.
In analogy with the functions that help the user comply with the criteria for creating the appropriate weighted edgelist, there is also a function that transforms longitudinal data into the format outlined above. The as.longitudinal-function does two operations. First, it adds the missing columns if either the fourth and fifth or just the fifth column are missing in the dataset, e.g. if only the time, the creator, and target are known. If this is the case, it assumes that all ties carry a weight of 1 (fourth column), and that there are no isolates in the network. Second, it reorganises the identification numbers so that they follow a strict sequence. If the identification numbers of nodes were not entered in a order strict, the function will reassign the identification numbers and give a warning message. A table containing the association between the original and the new identification numbers will be attached to the data object as an attribute called order. This table is used by the other functions in tnet to rearrange node attribute data when this type of data is included in the analysis. Table 11 shows how the as.longitudinal-function works on an object where the fourth and fifth column are missing and the identification numbers are not in sequence.
>data 2008-09-12 13:45:00 1 5 2008-09-12 13:46:31 4 2 2008-09-12 13:49:27 1 2 >transformed.data <- as.longitudinal(data) >transformed.data 2008-09-12 13:45:00 1 2 1 2 2008-09-12 13:46:31 3 4 1 4 2008-09-12 13:49:27 1 4 1 4 >attributes(transformed.data)$order 1 1 5 2 4 3 2 4
Table 11: How the as.longitudinal-function works on an object called data. Lines starting with > refer to commands in R. The first command displays the object. This object is then the input of the as.longitudinal-function. The third command displays this output of the function, transformed.data. The last command displays the attribute order, which is the key between the old and new node identification numbers.
A main issue with longitudinal network datasets is that the severing of ties is often not recorded. For example, we know that an email was sent, but we do not know when the tie between the sender and receiver is broken. There are several ways of estimating the severing of ties, such as the introduction of a smoothing window (e.g. Kossinets and Watts, 2006; Panzarasa et al., 2009). This method assumes that social ties are severed if there has been a prolonged period of time with no interaction (i.e. the length of the window). A function that allows for the severing of ties after a set amount of time and adds negative ties (i.e., where the fourth column is -1) in longitudinal datasets is add_window_to_longitudinal_data.
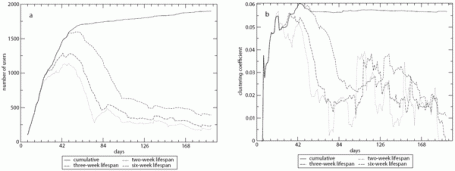
Since longitudinal datasets are richer in detail than static weighted ones, the function longitudinal_data_to_edgelist was created to transform a longitudinal network into a static network. This function is particularly useful when studying the evolution of static network measures over time (e.g. Kossinets and Watts, 2006; Panzarasa et al., 2009). A possible application of add_window_to_longitudinal_data and longitudinal_data_to_edgelist-functions to the Online Social Network (see Chapter 1) is illustrated in Figure 18. The Figure shows both cumulative networks (i.e., ties are never severed) and networks constructed with smoothing windows of 2, 3, and 6 weeks. Both panels in Figure 18 highlight the vulnerability of network measures to the use of a smoothing window. Panel a suggests that there is only a small core of users that actively use the virtual community at the end of the observation period. An analysis of the cumulative network at that point would be heavily influenced by the majority of users that only used the network in the first 6 weeks, and would not reflect the current activities that are occurring in the community. This could bias network measures and, ultimately, the analysis. Panel b shows the evolution of one possible measure, the clustering coefficient (Chapter 2). In particular, the clustering coefficient measured on the active core fluctuates greatly and is mostly below the value found in the cumulative network.

Trackback this post | Subscribe to the comments via RSS Feed