Random Networks
When analysing network and computing measures, it is difficult to gauge whether the outcome is “high” or “low”. For example, is a clustering coefficient of 0.3 high or low? Random networks based on an observed network can be used as to create a benchmark value. In other words, an observed value (e.g., a clustering coefficient) can be compared to the distribution of clustering coefficient from corresponding random networks.
There are many procedures or null models for creating random networks. These procedures vary in how much they randomise an observed network. Traditionally, classical (also known as Bernoulli or Erdos-Reyni) random networks were the commonly used. These networks are constructed by using the same number of nodes, and assigning each dyad a uniform probability of having a tie based on the number of observed ties. In these networks, the degree distribution is uniform or Poisson. However, such a distribution rarely exist in reality. In fact, most real-world distributions are skewed.
In response to the lack of comparability between observed networks and classical random networks, a set of randomisation procedures that maintain additional features has been developed. First, by reshuffling ties in an observed network, it is possible to create random networks that maintain each node’s number of ties as well as the overall number of nodes and ties (Molloy and Reed, 1995). Second, for weighted networks, it is also possible to reshuffle only the weights. By maintaining the ties and simply reshuffling tie weights, features based on connectedness are maintained (e.g., the size of the largest interconnected group of nodes; Opsahl et al., 2008).
Classical Random Networks
Classical random networks are created by setting the number of nodes and giving all possible ties a uniform probability of being formed. Given that each tie is independent of all other ties, it is often possible to approximate the value of a measure mathematically without the need of simulations when using these networks. For example, the global clustering coefficient is equal to the uniform probability and the average shortest path is approximately the ratio between the logarithm of nodes and the logarithm of the average number of ties that nodes have (Watts and Strogatz, 1998). A main limitation of classical random networks is that the distribution of ties across nodes is uniform or Poisson. While this in convinient for deriving expected values, such a distribution rarely exist in reality. In fact, most real-world degree distributions are skewed, which implies that a few nodes are hubs. The hubs particulily affect network measures based on shortest paths as hubs reducing it by acting as shortcuts among other nodes.
Link Reshuffling
As nodes’ degree are not uniformely distributed, a second procedure for creating random networks consists in reshuffling the topology, reaching the maximally random network with the same degree distribution as the observed network (Maslov and Sneppen, 2002; Newman, 2003). It does so by randomly selecting two ties, and
. The two ties are then rewired by setting
and
. The weights are automatically redistributed by remaining attached to the reshuffled ties. However, if either of these ties is already formed, this step is reverted, and two new ties are selected. This condition guarantees that multiple ties are not formed between two nodes, which ensures that the weight and degree distributions remain unchanged. If this procedure is repeated enough times, the outcome is a corresponding random network. This model is commonly used in the Physics literature; however, each of the random networks that are produced is not produced with an equal probability. For more details, see Snijders (2001) and Rao et al. (1996).
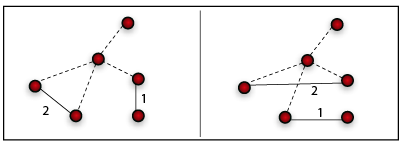
One step of the link reshuffling procedure for an undirected netwok. Note that while a node maintain its out-strength in a directed network, this is not the case in an undirected network.
Weight Reshuffling
While the link reshuffling procedure can be applied to binary networks, two other procedures are possible to use when dealing with weighted networks. The weight reshuffling procedure consists simply in reshuffling the weights globally in the network (Opsahl et al., 2008). This null model maintains the topology of the observed network. Therefore, the number of ties originating from a node (degree) does not change. Moreover, other features, such as the giant component does not change as well.
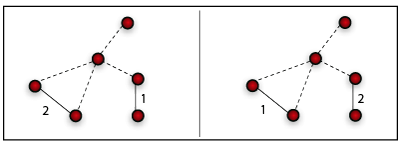
One step of the weight reshuffling procedure for an undirected netwok
Local Weight Reshuffling
Inevitably, since weights are reshuffled globally using the link and weight reshuffling procedures, they produce random networks in which the nodes do not maintain the same node strength as in the observed network. Local weight reshuffling is a third randomisation procedure for directed networks that preserves this quantity by reshuffling weights locally for each node across its outgoing ties (Opsahl et al., 2008). The procedure can be extended to undirected networks by duplicating an undirected tie into two directed ties – one in each direction. It should be noted that this procedure breaks the weight symmetry in the two directions of an undirected tie (the topology remains invariant). The appropriateness of this method for undirected networks depends on the research setting and how tie weights are defined. For example, its applicability to undirected transportation networks is justified by the typically directed nature of traffic flows (although the US airport network displays a high symmetry; Barrat et al., 2004).

One step of the local weight reshuffling procedure for a directed netwok
Example
To highlight how the various procedures differ, the diagram below shows an observed network and the four random networks introduced above. To reproduce these networks, see the R code below.
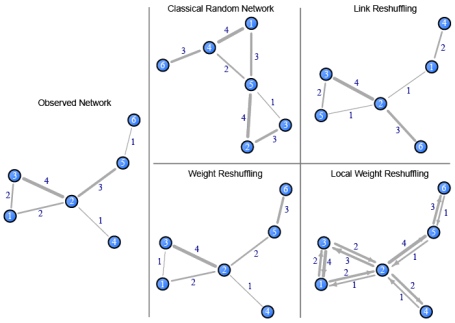
Random networks created using the null models outlined above. The classical random network uses the same number of nodes, a 0.4 uniform probability of ties being present, and tie weights randomly picked between 1 and 4.
Want to test it with your data?
The randomisation procedures are implemented in tnet. First, you need to download and install tnet in R. Then, you need to create an edgelist of your network (see data structures in tnet for weighted one-mode networks). The commands below show how the edgelist for the sample network here can manually be entered, and how to apply the randomisation procedures.
# Load tnet library(tnet) # Create a Classical Random Network with 6 nodes and a 0.4 uniform probability of ties being created rg_w(nodes = 6, arcs = 0.4, weights = 1:4, directed = FALSE) # Load an observed network net <- cbind( i=c(1,1,2,2,2,2,3,3,4,5,5,6), j=c(2,3,1,3,4,5,1,2,2,2,6,5), w=c(2,2,2,4,1,3,2,4,1,3,1,1)) # Link reshuffling of the sample network rg_reshuffling_w(net, option="links") # Weight reshuffling of the sample network rg_reshuffling_w(net, option="weights") # Local weight reshuffling of the sample network rg_reshuffling_w(net, option="weights.local")
To recreate the illustration of the random networks, the code below is the first step. The diagram above was made a bit nicer afterwards using Adobe Illustrator.
# Load tnet and the observed network library(tnet) net <- cbind( i=c(1,1,2,2,2,2,3,3,4,5,5,6), j=c(2,3,1,3,4,5,1,2,2,2,6,5), w=c(2,2,2,4,1,3,2,4,1,3,1,1)) # Create object to hold the random networks rn <- list() # Put the four random networks in the object (using seed=1 for replication purposes) rn[[1]] <- rg_w(nodes = 6, arcs = 0.4, weights = 1:4, directed = FALSE, seed = 1) rn[[2]] <- rg_reshuffling_w(net, option="links", seed=1) rn[[3]] <- rg_reshuffling_w(net, option="weights", seed=1) rn[[4]] <- rg_reshuffling_w(net, option="weights.local", seed=1) # Convert networks to igraph format net <- tnet_igraph(net) rn <- lapply(rn, function(a) tnet_igraph(a)) # Set up plot par(mfrow=c(2,3), pty="m", mar=c(4,5,0.1,0.1), xaxs="r", yaxs="i") # Get layout of observed network layout <- layout.fruchterman.reingold(net) # Plot observed network plot(net, main="Observed Network", layout=layout, vertex.label=1:6, edge.width=E(net)$weight, edge.label=E(net)$weight) # Plot random networks plot(rn[[1]], main="Classical Random Network", layout=layout.fruchterman.reingold, vertex.label=1:6, edge.width=E(rn[[1]])$weight, edge.label=E(rn[[1]])$weight) plot(rn[[2]], main="Link Reshuffling", layout=layout.fruchterman.reingold, vertex.label=1:6, edge.width=E(rn[[2]])$weight, edge.label=E(rn[[2]])$weight) plot.new() plot(rn[[3]], main="Weight Reshuffling", layout=layout, vertex.label=1:6, edge.width=E(rn[[3]])$weight, edge.label=E(rn[[3]])$weight) plot(rn[[4]], main="Local Weight Reshuffling", layout=layout, vertex.label=1:6, edge.width=E(rn[[4]])$weight, edge.label=E(rn[[4]])$weight, edge.arrow.size=0.5)
References
Barrat, A., Barthelemy, M., Pastor-Satorras, R., Vespignani, A., 2004. The architecture of complex weighted networks. Proceedings of the National Academy of Sciences 101 (11), 3747-3752. arXiv:cond-mat/0311416
Molloy, M., Reed, B., 1995. A critical point for random graphs with a given degree sequence. Random Structures and Algorithms 6, 161-180.
Opsahl, T., Colizza, V., Panzarasa, P., Ramasco, J. J., 2008. Prominence and control: The weighted rich-club effect. Physical Review Letters 101 (168702). arXiv:0804.0417.
Watts, D. J., Strogatz, S. H., 1998. Collective dynamics of “small-world” networks. Nature 393, 440-442.
1. Jim | January 16, 2012 at 12:19 pm
Jim | January 16, 2012 at 12:19 pm
Hi Tore, great site and resource.
I wondered how well the algorithm scaled to larger networks? I have a network of over 10k nodes and about 80k edges, I have been running your program for a while so far, but wondered if you had any idea on how long it might take to calculate?
Many thanks, and again, great site.
2. Tore Opsahl | January 16, 2012 at 4:28 pm
Tore Opsahl | January 16, 2012 at 4:28 pm
Hi Jim,
Glad you are finding it useful!
The link reshuffling is not very efficient as it relies on igraph. I used to have a much more efficient script for sparse graphs, but had to provide loads of support for people using it on dense graphs. If you send me an email, we can see if this might be more appropriate for you.
Best,
Tore
3. Marton | November 10, 2014 at 6:28 am
Marton | November 10, 2014 at 6:28 am
Hi Tore,
Many thanks for your site, I find it very helpful!
I have a question regarding the random network. How should I compare the metrics in the observed network with those in random network? I suppose I should use a kind of significance test (like in the weighted rich-club effect), but how can I implement that in R? Or doesn’t it need significance test? When can I say my outcome is high or low?
Sorry for the trivial question, but I’m just a beginner in the network analysis
Thanks,
Marton
4. Tore Opsahl | November 11, 2014 at 2:08 am
Tore Opsahl | November 11, 2014 at 2:08 am
Hi Marton,
It’s great that you are thinking along these lines. I would recommend the following:
– calculate the metric of interest for the actual network (“observed”)
– in a loop of at least 1,000
{
– create a random network
– calculate the metric on the random network
}
– compare the observed value to the ones found using random networks
If the observed value is in the far tails of the distribution of values from random networks, then you can make a claim about it being different from random.
I would recommend using the reshuffling procedures for generating random network to ensure the network are comparable.
Good luck,
Tore
5. Marton | November 11, 2014 at 2:43 pm
Marton | November 11, 2014 at 2:43 pm
Thanks Tore, but what does it mean “loop” here? I suppose it’s something that makes me 1000 random networks, but how can I implement it? Can you give me an example command to implement this comparision? Sorry again for my ignorance!
6. Tore Opsahl | November 11, 2014 at 4:32 pm
Tore Opsahl | November 11, 2014 at 4:32 pm
No worries, Marton. There are many who have the same question, so here is an example.
Research question: Are closed triplets composed of stronger ties than we would expect by chance in the c.elegans neural network?
Method: Compare the global clustering coefficient of the c.elegans network to the one found in comparable network where the weights have been reshuffled.
Code:
# Load tnet
library(tnet)
# Load tnet datasets
data(tnet)
# Pick the celegans as the network of interest
netObs <- celegans.n306.net
# Clean up
rm(list=ls()[ls() != "netObs"])
# Compute the global clustering coefficient
valueObs <- clustering_w(netObs, measure="am")
valueObs
# 0.2364436
# Let’s generate random networks!
# Output vector
valueRdm <- rep(NaN, 1000)
# Loop
for(i in 1:length(valueRdm)) {
# Use weight reshuffling to generate comparable networks with seed so results are reproducable
netRdm <- rg_reshuffling_w(netObs, option="weights", seed=i)
# Calculate metric on random network
valueRdm[i] <- clustering_w(netRdm, measure="am")
}
# Plot results (or do other tests)
xaxisRange <- c(min(c(valueRdm, valueObs-.01)), max(c(valueRdm, valueObs+.01)))
hist(valueRdm, xlim=xaxisRange)
# Add vertical line for observed value
abline(v=valueObs, col="red", lwd=3)
Results:

As you can see, the observed value (vertical red line) is much higher than the comparable random values. You could also compute the standard deviations of valueRdm to determine the statistical significance of this.
7. Marton | November 14, 2014 at 7:45 am
Marton | November 14, 2014 at 7:45 am
Many thanks Tore, I’m really gratful for your help!
8. Marton | November 16, 2015 at 3:07 pm
Marton | November 16, 2015 at 3:07 pm
Hi Tore,
Now I have some problem with the “rg_reshuffling_w” function. So far I have used this function with the link reshuffling method and it workd properly. But now I can’t used this; when I run the command I get the error message: “Error in igraph::rewire(net.i, niter = (ecount(net.i) * 10)) : unused argument (niter = (ecount(net.i) * 10))”. After this I run those scripts that I used with this function before and I got the same error message again (they had worked properly back then I had worked with them). I tried the links option with the example given in the offical tnet package description but it doesn’t work either. Do you have any idea what happened with this function? Was there any change with it? Or maybe am I doing something wrong?
Thanks,
Marton
9. Tore Opsahl | November 17, 2015 at 11:51 pm
Tore Opsahl | November 17, 2015 at 11:51 pm
Hi Marton,
Thanks for spotting an error! It seems that igraph has been updated and the tnet code that connected to it has broken. I will look into this and upload a new version of tnet to Cran when I’ve fixed it.
Best,
Tore
10. Tore Opsahl | November 18, 2015 at 3:59 am
Tore Opsahl | November 18, 2015 at 3:59 am
Hi Marton: An updated version has been created, and a new version of tnet (3.0.14) has been uploaded to CRAN. If you run, update.packages(), you should see tnet being updated. It might take some time before tnet gets updated on all the mirrors. Good luck! Tore
11. Marton | November 20, 2015 at 9:03 am
Marton | November 20, 2015 at 9:03 am
Hi Tore,
it works! Thank you for your update!
best,
Marton
12. Arlo | March 22, 2021 at 6:03 pm
Arlo | March 22, 2021 at 6:03 pm
Hi Tore,
Thanks for your great and inspiring work! I have a question for local weight reshuffling. If the random graph created by this method is directed, can I use it to compare parameters like weighted clustering coefficient and weight average path length with the observed network (weighted but un-directed)? Thanks!
13. Tore Opsahl | March 23, 2021 at 3:43 pm
Tore Opsahl | March 23, 2021 at 3:43 pm
Hi Arlo,
I think then you are not exactly comparing apples to apples then. Depends on your setting, but generally I would recommend creating undirected random networks if your network is undirected.