Import and Export Functions
Networks can be imported and exported into tnet from a range of other network analysis programmes. This page refers to weighted one-mode networks as most programmes cannot deal with two-mode and longitudinal networks.
UCINET
UCINET is a Windows software for general analysis of social network data (UCINET’s website). This programme is not free, but there is a 30-day trial period.
Import into tnet
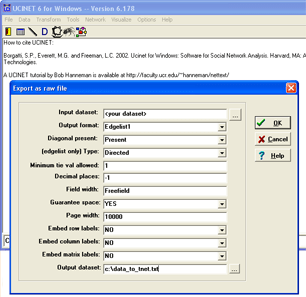 Weighted one-mode networks in UCINET can be analysed in tnet by following these steps. First, go to UCINET, and select Data > Export > Raw …
Weighted one-mode networks in UCINET can be analysed in tnet by following these steps. First, go to UCINET, and select Data > Export > Raw …
- Input dataset: <your dataset>
- Output format: Edgelist1
- (edgelist only) Type: directed
- Minimum tie val allowed: 1
- Embed row labels: NO
- Embed column labels: NO
- Output dataset: c:\data_to_tnet.txt
(this can be changed, but then you need to changed the entries further down in this example)
Second, an exclamation mark at the end of the outputted file must be manually removed. UCINET adds an exclamation mark at the end of each file, and this cannot be read by R/tnet in a simple way. One way of doing this, is to open c:\data_to_tnet in Notepad, and delete the last line (this should just be !).
Third, the file can be loaded in R as an object called net by typing the following command (note that the \ has become /):
data <- read.table(file="c:/data_to_tnet.txt")
Export from tnet
There is a specific function in tnet for exporting networks to UCINET. This function is called tnet_ucinet. This function works by taking three parameters:
- net: This should be the network to export
- type: Which type of network this is (default is to auto-detect, see
as.tnet) - file: The desired file name of the network (default is to create a file in the working directory with the current time, for example, tnet_ucinet_network-2011-04-10_150817.dl).
The file can then be loaded in UCINET by using the DL import function. You can find this function through the menu: Data > Import > DL. When the function’s dialog box opens, you must select the downloaded file containing the dataset by clicking on “…” after “Input text file in DL format”. The second box can be set to the default, but do remember, and change if you wish, the name that appears in the third box as this will be the name of the internal UCINET file.
The R-package igraph
igraph is a free software package for creating and manipulating undirected and directed graphs. This package has been implemented in R, and tnet depends on it for various heavy calculations.
Import into tnet
As both igraph and tnet are implemented in R, data can easily be transferred between the two packages. The following code shows how to extract the edgelist from igraph, and use it in tnet.
# Load igraph and tnet library(tnet) # Create a sample igraph network with tie weights g <- erdos.renyi.game(6, 0.4, type="gnp") E(g)$weight <- sample(1:4, size=ecount(g), replace=TRUE) # Extract edgelist and a 1 to the node id's as igraph starts from 0 while tnet 1. net <- cbind(get.edgelist(g, names=FALSE), E(g)$weight) # For binary networks, a third column of 1s must be added if(ncol(net)==2) net <- cbind(net, 1) # For undirected networks, the symmetrise function must be run if(!is.directed(g)) net <- symmetrise_w(net) # Ensure that it conforms to the tnet standard net <- as.tnet(net, type="weighted one-mode tnet")
Export from tnet
There is a special function in tnet for exporting networks to igraph (tnet_igraph). Below is an example of how it can be used.
# Load igraph and tnet library(tnet) # Create a sample tnet network net <- rg_w(nodes=10, arcs=0.4, weights=1:4, directed=FALSE) # Create igraph object g <- tnet_igraph(net,type="weighted one-mode tnet") # Get summary statistics summary(g)
Pajek
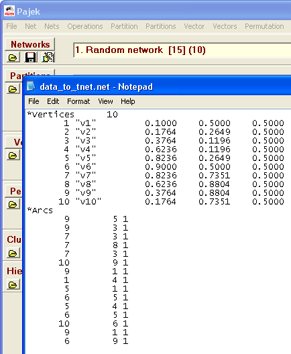 Pajek is a Windows programme for analysis and visualization of large networks.
Pajek is a Windows programme for analysis and visualization of large networks.
Import into tnet
To export your data from Pajek to tnet, you must first save the network in a Pajek .net file, you can do this by going to File > Network > Save, and selecting a location. In this example, I will use c:\data_to_tnet.net. Then you need to open the .net file, e.g. c:\data_to_tnet.net in Notepad as shown on picture to the right. You need to delete the lines from “*Vertices” to, and including, “*Arcs”, so that all you have left is three columns with numbers. Then save the file.
Then, you can open R, and type (note that the \ has become /):
data <- read.table(file="c:/data_to_tnet.net")
The R-package sna
The sna R-package is a collection of routines for social network analysis. It has been developed by Carter Butts at University of California, Irvine.
Import into tnet
To import an sna-object into tnet, you must use the as.edgelist.sna-function.
# Load sna and tnet library(sna) library(tnet) # Create a sample sna network g <- rgraph(n=10, tprob=0.4, mode="graph") # Extract edgelist net <- as.edgelist.sna(g) # Ensure that it conforms to the tnet standard net <- as.tnet(net, type="weighted one-mode tnet")
Export from tnet
To load a network into sna, a matrix must be created. This can be done using the following code:
# Load sna and tnet
library(sna)
library(tnet)
# Create a sample tnet network
net <- rg_w(nodes=6, arcs=0.4, weights=1:4, directed=FALSE)
# Create sna object
net <- as.matrix(net)
N <- max(c(net[,"i"],net[,"j"]))
g <- matrix(data=0, nrow=N, ncol=N)
g[net[,c("i","j")]] <- net[,"w"]
# Ensure that it conforms to the sna standard
g <- as.sociomatrix.sna(g)
The R-package network
The R-package network is a basic package that many other R-packages use for their data handling. As this package cannot handle weighted data, they are remove in any transfer process.
Import into tnet
To import a network-object into tnet, the network can be extracted as an edgelist using the as.matrix.net with the matrix.type-parameter set to edgelist. Below is an example.
# Load sna and tnet library(network) library(tnet) # Create sample network object g <- network.initialize(6, directed=FALSE) g <- add.edge(g, 1, 2) g <- add.edge(g, 1, 3) g <- add.edge(g, 2, 4) g <- add.edge(g, 2, 5) g <- add.edge(g, 5, 6) # Extract edgelist, and add a column of 1's net <- as.matrix.network(g,matrix.type="edgelist") net <- cbind(net, 1) # For undirected networks, the symmetrise function must be run if(!network::is.directed(g)) net <- symmetrise_w(net) # Ensure that it conforms to the tnet standard net <- as.tnet(net, type="weighted one-mode tnet")
Export from tnet
To transfer a tnet object into the network package, …
# Load sna and tnet
library(network)
library(tnet)
# Create a sample tnet network
net <- rg_w(nodes=6, arcs=0.4, weights=1:4, directed=FALSE)
# Create sna object
net <- as.matrix(net)
N <- max(c(net[,"i"],net[,"j"]))
g <- matrix(data=0, nrow=N, ncol=N)
g[net[,c("i","j")]] <- net[,"w"]
g <- network(g, directed=FALSE)
1. ryan | June 14, 2016 at 4:59 pm
ryan | June 14, 2016 at 4:59 pm
Attempting to use the above code for igraph import and getting an error. Rerunning your code above:
Using: $version.string
[1] “R version 3.3.0 (2016-05-03)”
$nickname
[1] “Supposedly Educational”
>
> # Load igraph and tnet
> library(tnet)
>
> # Create a sample igraph network with tie weights
> g E(g)$weight
> # Extract edgelist and a 1 to the node id’s as igraph starts from 0 while tnet 1.
> net
> # For binary networks, a third column of 1s must be added
> if(ncol(net)==2)
+ net
> # For undirected networks, the symmetrise function must be run
> if(!is.directed(g))
+ net
> # Ensure that it conforms to the tnet standard
> net
> clustering_tm(net)
Error in clustering_tm(net) : Network not loaded properly
2. Tore Opsahl | June 14, 2016 at 5:36 pm
Tore Opsahl | June 14, 2016 at 5:36 pm
Hi Ryan,
It seems you are missing line #5 in the above example. This line creates a random igraph network. Let’s connect over email if you are struggling with your own data.
Tore
3. Misi | August 10, 2016 at 12:29 pm
Misi | August 10, 2016 at 12:29 pm
Hi Tore,
your tnet library is great! I have a problem importing data from igraph though. I followed your instructions above which work fine, but result in an edgelist which has vertex ranks as IDs rather than the vertex IDs I would like to use (numeric company IDs such as “5565877668”). I run the following starting from my “g2up” igraph data.frame:
g2uptn <- cbind(get.edgelist(g2up, names=TRUE), E(g2up)$weight)
if(!is.directed(g2up))
g2uptn <- symmetrise_w(g2uptn)
g2uptn <- as.tnet(g2uptn , type="weighted one-mode tnet")
which gives me the following error message: "Error in Ops.factor(net[, "i"], net[, "j"]) : level sets of factors are different"
Creating a data.frame with numeric IDs doesn't seem to solve the problem either. Running this code I run into the same error message:
g2uptn <- cbind(get.edgelist(g2up, names=TRUE), E(g2up)$weight)
colnames(g2uptn)<-c("i", "j", "w")
g2uptn2 <- data.frame(g2uptn , row.names = NULL)
if(!is.directed(g2uptn2 ))
g2uptn2 <- symmetrise_w(g2uptn2 )
g2uptn2 <- as.tnet(g2uptn2 , type="weighted one-mode tnet")
Any help would be very welcome!
Best,
Misi
4. Tore Opsahl | August 12, 2016 at 2:52 pm
Tore Opsahl | August 12, 2016 at 2:52 pm
Hi Misi,
Glad you are finding the library useful. tnet is written to handle networks with integer/numeric node identifiers. Based on your error messages, it seems that the data class for the identifiers is factor. Try to type class(g2uptn[,1]) to see the data class. Also, you might want to turn of names=TRUE in the igraph function get.edgelist.
Best,
Tore
5. Misi | August 15, 2016 at 3:39 pm
Misi | August 15, 2016 at 3:39 pm
Hi Tore,
great, this solved my initial problem, very sorry I missed such a basic problem. Now after running:
g2uptn <- cbind(get.edgelist(g2up, names=TRUE), E(g2up)$weight)
colnames(g2uptn)<-c("i", "j", "w")
g2uptn2 <- data.frame(g2uptn , row.names = NULL)
class(g2uptn2[,1])
[1] "factor"
g2uptn2$i <- as.numeric(as.character(g2uptn2$i))
g2uptn2$j <- as.numeric(as.character(g2uptn2$j))
g2uptn2$w <- as.numeric(as.character(g2uptn2$w))
class(g2uptn2[,1])
[1] "numeric"
g2uptn2 <- symmetrise_w(g2uptn2)
I get the following new error message:
Error in as.tnet(net, type = "weighted one-mode tnet") :
Not all node id's are integers
In addition: Warning message:
In as.tnet(net, type = "weighted one-mode tnet") :
NAs introduced by coercion to integer range
Now, this is surprising as I do think that all my IDs are integers and there are no NAs, I manually checked all 3000 of them. This is my first few lines:
i j w
1 5562859370 5560510272 1
2 5562859370 5567267488 2
3 5562859370 5567066088 1
4 5562859370 5567980031 1
5 5562859370 5561400283 1
6 5562859370 5564857984 1
7 5562859370 5567137962 1
8 5562859370 5566225347 1
9 5562859370 5566862503 1
10 5562859370 5561852103 1
Any ideas on what may cause this new error?
Best,
Misi
6. Tore Opsahl | August 19, 2016 at 12:46 pm
Tore Opsahl | August 19, 2016 at 12:46 pm
Hi Misi,
I am not entirely sure what’s is happening. If you send me an email with your data and code, I will have a look at it.
Tore